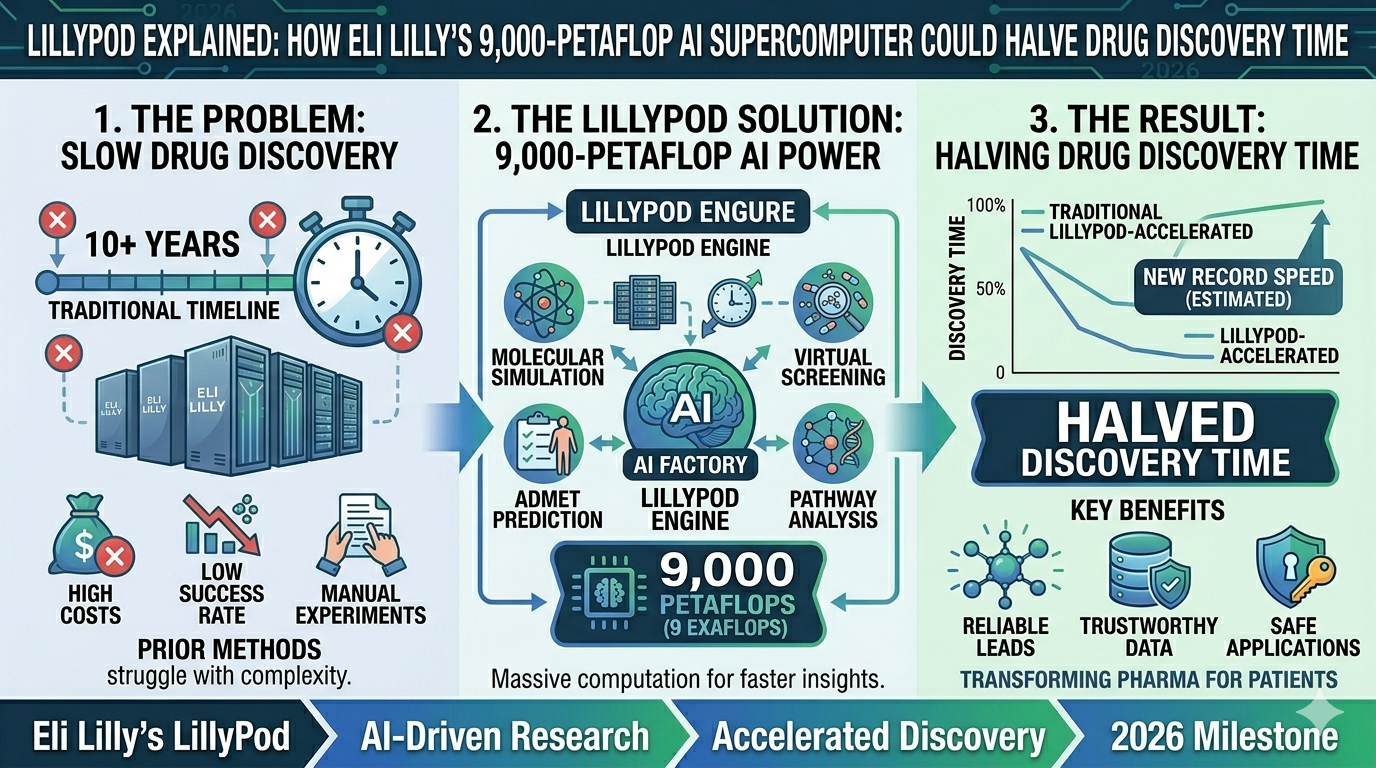
Drug discovery has always been a waiting game measured in decades. A promising molecule enters the lab. It gets tested. It fails. Another molecule enters. Repeat — thousands of times, across years of experiments, at costs that can exceed $2 billion before a single drug reaches a patient. The fundamental constraint has always been the wet lab: scientists can only run so many physical experiments per year, testing roughly 2,000 molecular ideas per target before resources run out.
On February 26, 2026, Eli Lilly inaugurated LillyPod and changed that constraint. Powered by a DGX SuperPOD with 1,016 NVIDIA Blackwell Ultra GPUs, Lilly's AI factory delivers more than 9,000 petaflops of AI performance. It was assembled in just four months. The supercomputer creates what Lilly's leadership calls a "computational dry lab" — a virtual research environment where scientists can now test billions of molecular hypotheses in parallel, at their fingertips, before committing a single experiment to the physical world.
From 2,000 Molecules to Billions: What LillyPod Changes
The numbers that matter most from LillyPod's launch are not the GPU count or the petaflop rating. They are the comparison between what was possible before and what is possible now.
Historically, drug discovery teams have been limited to analyzing roughly 2,000 molecular ideas per target per year due to wet-lab constraints. "Now the supercomputer center essentially just breaks the physical limit of the wet lab," said Yue Wang Webster, vice president of research and development informatics at Lilly. "Now in the dry lab, you can test billions of molecule ideas at your fingertips."
This is not an incremental improvement. Moving from 2,000 to billions of parallel molecular simulations per year is a qualitative shift in how pharmaceutical science can be conducted. It changes which questions can even be asked — because asking a question that requires testing 10 million molecular configurations was previously impossible. Now it is a morning's compute run.
It's a big day for us with the supercomputer coming on board, but it's a day 150 years in the making. LillyPod is a powerful symbol of who we are and why we do this work: to make life better for people around the world.
Diogo Rau, Executive Vice President and Chief Information and Digital Officer, Eli Lilly, February 26, 2026What LillyPod Is Actually Being Used For
Workloads on LillyPod on day one span genomics, peptide design, single-cell biology, imaging, and manufacturing. This breadth reflects Lilly's chief AI officer Thomas Fuchs's insistence that AI at Lilly is not a single-use discovery tool. It is embedded across the entire pharmaceutical value chain.
| Application Area | What LillyPod Enables | Impact |
|---|---|---|
| Genomics | Analysis of 700 TB of genomic data using 290+ TB of GPU memory | Faster identification of disease targets and patient stratification |
| Molecular Design | Simulation of billions of small-molecule and peptide candidates in parallel | Dramatically expands the chemical space explored per target |
| Single-Cell Biology | AI models trained on millions of single-cell experiments | Deeper understanding of disease mechanisms at cellular level |
| Clinical Development | AI-designed trials optimizing endpoints and patient populations | Reduces trial failures from poor trial design |
| Manufacturing | AI optimization of production processes and quality control | Reduces manufacturing variability and accelerates scale-up |
The Computing Scale Behind LillyPod
In 1992, Lilly's Cray supercomputer was the pinnacle of scientific computing. Today, a single NVIDIA Blackwell Ultra GPU in the new AI factory contains the power of approximately 7 million Cray systems. And since the AI factory comprises 1,016 Blackwell Ultra GPUs, it delivers over 9,000 petaflops of AI performance. This means it can do over 9 quintillion math problems every second.
The system's nearly 5,000 connections are built from more than 1,000 pounds of fiber cables. Lilly has committed to running the entire infrastructure on 100 percent renewable electricity by 2030, using efficient liquid cooling to minimize incremental energy impact. This sustainability commitment matters because LillyPod's scale of compute is not modest — maintaining it long-term requires a thoughtful energy strategy.
TuneLab: Sharing the Benefit with the Broader Biotech Ecosystem
LillyPod is not just an internal tool. Lilly is using it to build and share AI models through TuneLab, its federated AI platform for drug discovery. A portion of LillyPod's AI models will be accessible through TuneLab, the company's federated AI platform for early-stage drug discovery, trained on over $1 billion worth of research data.
TuneLab uses federated learning to allow smaller biotech companies to access Lilly's models while keeping their own proprietary data private and isolated. This means even companies without LillyPod-scale infrastructure can benefit from models trained on Lilly's 150 years of pharmaceutical data — including, as Fuchs noted, lessons learned from millions of molecules that failed.
The bottom line for healthcare and patients: LillyPod does not guarantee faster cures. Drug discovery still requires clinical trials, regulatory approval, and manufacturing scale-up. But by radically compressing the earliest stages of the process — target identification, molecule design, and experimental hypothesis generation — LillyPod can meaningfully shorten the overall timeline. If the traditional 10-year pathway from first human dose to market launch can be compressed even by 20 to 30 percent, the human impact across thousands of disease programs globally is enormous.